Cells are able to transmit and process information by using molecular machines designed by evolution. These biochemical machines, usually proteins, change their structure and function in predictable ways based on input from the environment. These components of the cell can be organized into cascades, or branching chain reactions, in which the changes in one component are transmitted to a large number of components downstream.
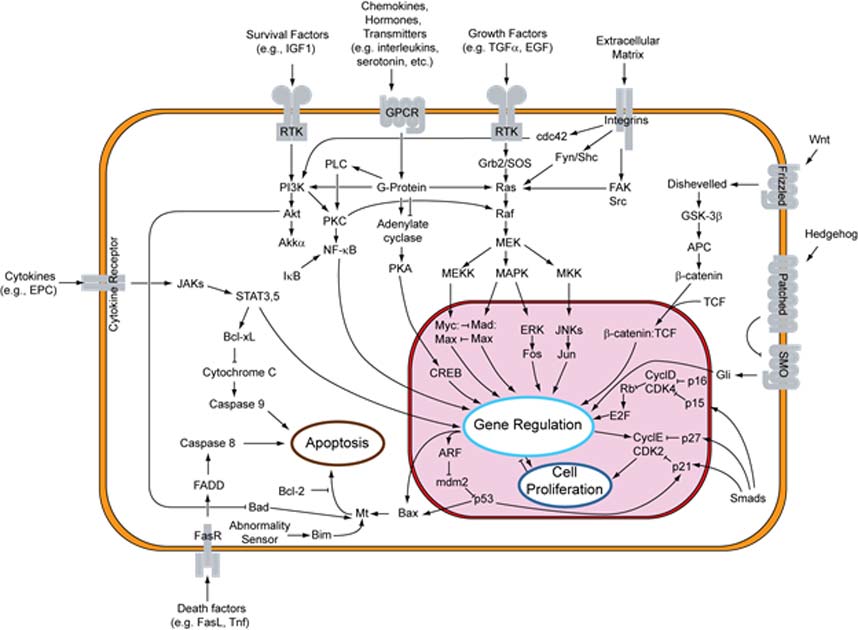
Signal transduction pathway. (CC BY-SA 3.0)
In a recent paper by Dr. Jie Song and colleagues based at Emory University, they report on a molecular machine composed of synthesized DNA that incorporates some of the traits of a biological system.
Protein Machines
Protein machines work in a cell because they have more than one stable structure. The structures are related and can convert back and forth between each other. However, one is more stable than the others in a given context because it is stabilized by input from the surrounding environment, perhaps by another protein that functions upstream in a sequence of reactions.
DNA Machine Parts
An antijunction consists of a DNA structure made of four double strands of equal length. They form a rectangle or square shape. Song and colleagues designed several kinds of antijunctions with more than one stable structure that can interconvert. The two different versions were arbitrarily dubbed ‘red’ and ‘green’.
In the version pictured below, a blue scaffold strand forms the foundation of the rectangle. Green, magenta, purple, and tan strands help form four double helices. The black ‘x’ marks the ends of the strands.
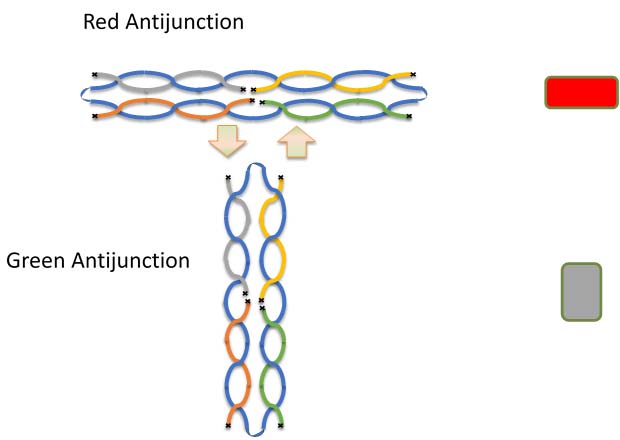
These structures are quite different from ‘junctions’ that are observed in other contexts. In junctions, double stranded DNA strands seem to cross over each other. Pictured here:
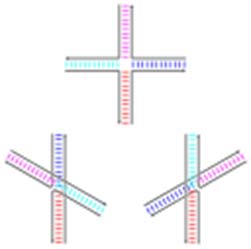
Schematic diagrams of the three base-stacking conformational isomers of the Holliday junction. By Antony-22 (Own work) (CC BY-SA 4.0)
The difference between the red and green antijunctions is subtle. Notice that in the red structure the green and magenta strands face each other, while in the green structure the green strand faces the purple.
Stacking in DNA
To understand the significance of this change, one needs to understand DNA structure.
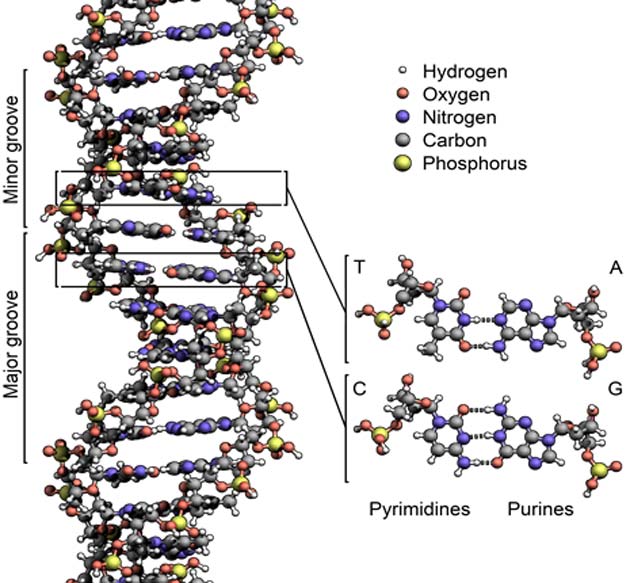
The structure of the DNA double helix. By Zephyris - Own work (CC BY-SA 3.0)
You may be familiar with the idea that the rings in DNA stick to each other in a specific pattern: T (thymine) and A (adenine) form one pair, while G (guanine) and cytosine (C) form the other. However, did you know that in this example the T-A pair is attracted to the C-G pair? In fact, any DNA pair is attracted to the pairs above and below it. This is called ‘stacking’, as if the DNA pairs were plates stacked on top of each other in the middle of the molecule. Stacking results from electron clouds above and below the ring structures of G, A, T, and C. The electron clouds of one DNA pair overlaps with the clouds above and below it.

So, the red antijunction is stabilized by the stacking between the green/blue helix and the purple/blue helix as well as the magenta/blue and the tan/blue helices. In effect, the two helices form one big long helix on each side of the antijunction. In the green antijunction, the helices switch stacking partners. So, each antijunction structure is expected to be roughly as stable as the other, because they exhibit equal amounts of stacking. There is every reason to expect that they might interconvert given the right encouragement. Song and colleagues were able to use molecular triggers to induce the conversion from one antijunction form to the other.
Arrays Built of Antijunctions
In addition they were able to build the antijunctions into rows and columns, forming arrays. The interesting thing about these arrays, is that when one antijunction makes the conversion from red to green it induces nearby antijunctions to follow suit.
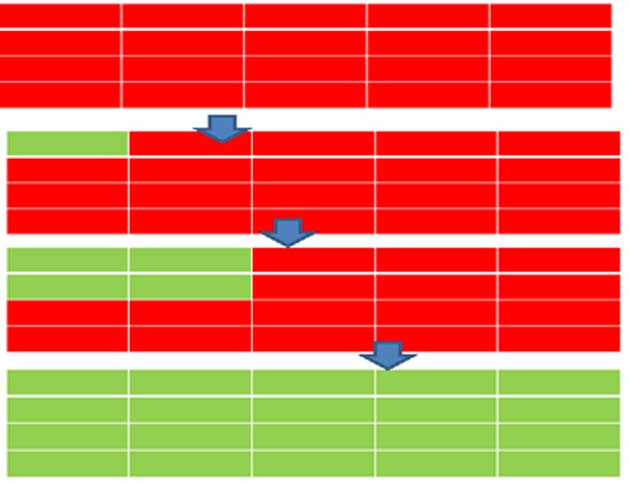
This is analogous to the activation of one protein by a structural change leading to activation by structural change in other proteins. This is the first time this kind of sustained chain reaction has been achieved in nanotechnology. Earlier molecular machines had one or a few moving parts. Much of the structure remained unchanged in the first molecular machines. In the machine built by Song and colleagues each component takes part in the transformation.
The conversion process can also be controlled. The starting point in the array can be selected by selecting where to put the triggers. The progress through the array can be directed by locking antijunctions into their original conformation. Antijunctions can then be unlocked at a later time. Because the green antijunction is elongated in one dimension compared to the red antijunction, arrays with corners can produce swinging arm-like movements.
Looking Towards the Future
The authors speculated on work they would like to do in the future. One limitation is the relatively slow speeds at which these transformations take place. Biological molecules cannot handle the high temperatures that are typically required for quick-moving industrial reactions. This is one area that needs improvement.
Finally, Song and colleagues are looking forward to producing arrays in “3D spaces, larger sizes, more intricately shaped designs, and more complex dynamic behaviors.”
Top image: Photo of the plastic grating on Watson’s book DNA (CC BY 2.0)
References
Eastman, Q., 2017. Switchable DNA mini-machines relay information. [Online]
Available at: http://news.emory.edu/stories/2017/06/ke_science_DNA_machines/
[Accessed 5 August 2017].
Masterson, A., 2017. DNA nanomachines can transmit information. [Online]
Available at: https://cosmosmagazine.com/technology/dna-nanomachines-can-transmit-information
[Accessed 5 August 2017].
Song, J. et al., 2017. Reconfiguration of DNA molecular arrays driven by information relay. Science Magazine, 357(6349), pp. 371-378.




No comment