Large-scale data storage is one of the most important forms of innovation that supports the abilities and potential of modern computing. It enables online functions and services such as Google and Amazon to function. In addition, the ability to keep high-volume intact and readable make forms of research such as genetic studies better and more effective.
On the flip side, however, masses of information that we’ve become accustomed to producing need extensive amounts of hardware and energy to maintain uncorrupted data and accessibility. This reason may be why researchers are always looking for new, more efficient means of data storage.
DNA is an example of potential new standards for the encoding and maintenance of information. It may seem surprising, but as DNA is nature’s data-coding and transcription solution, it makes more sense if humans harness this option.
The latest foray into the use of DNA to store data has been conducted, as part of a project, at the Waterford Institute of Technology.
Writing Data to DNA
The team behind it – including researchers from the University of Padua in Italy, such as lead author Federico Tavella – demonstrated the ability to encode human-language characters in DNA sequences, which is then stored in bacteria.
These microorganisms were a strain (or sub-type) of E. coli called Nova Blue. This modality of ‘writing’ data to DNA for storage is not unknown to science. In fact, the WIT/Padua team were building on previous work conducted in a similar vein.
However, the team also decided to proceed to a further stage in their project: solving the problem of how to ‘retrieve’ the data from its DNA ‘archive’ again.
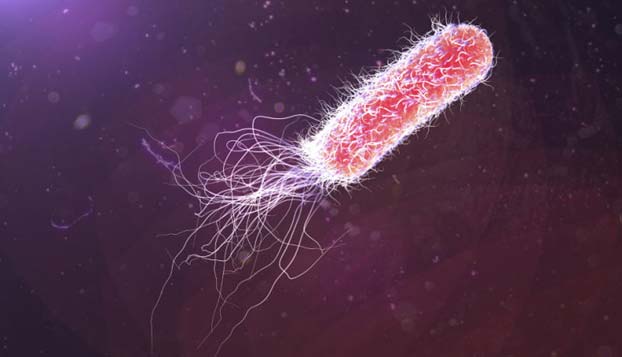
E. coli have the capacity to store data in the form of DNA sequences. (Source: Shutterstock)
The team started by converting their ‘data files’ to forms of DNA called plasmids. These are common features of bacteria, which use plasmids to store the sequences of potentially important proteins such as those involved in antibiotic resistance. Plasmids store this information remotely from the main bacterial genome for a variety of reasons, which include the rapid replication of plasmid proteins. One bacterial cell can also transfer one of its plasmids to another as a form of communication.
The researchers exploited this behavior to induce a Nova Blue cell to pick up one of their encoded plasmids and transfer it to a cell of another E. coli strain, HB101. The HB101 cell then gravitated towards a device that could translate the plasmid’s data into normal text again.
Accordingly, the team reported that the information carried in their complete prototype plasmid was a message reading ‘Hello World.'
Why Store Data in Bacteria?
The team published this success in DNA-data reading on arxiv.org, the Cornell University’s system for research archiving. It is an interesting description of how data stored on DNA can be recalled and converted into a usable format, rather than simply being archived.
A complete system of DNA as a medium for data has a number of theoretical advantages. DNA is, obviously, a well-validated method of information storage that can be transcribed, into different forms, with high fidelity and reliability. In addition, millions of DNA subunits, which could act as bits, can fit into the space taken up by a single cell. Therefore, this biological system can be capable of impressive volume and density, if scaled up.
Conversely, the data stored and moved around on the servers of today (which is estimated in terms of zettabyte or a billion bytes) may be able to fit in a much smaller form factor, if converted to a DNA-based standard.
But maintaining archives in such a biological format also implies drawbacks, some of which could be significant. DNA replication can be subject to mistakes, with a probability of being influenced by the organism or cell type. In real-world terms, this concern may translate to data loss or corruption.
Additionally, DNA data is read as a result of HB101 reaching the device, which enabled the researchers to do so. This behavior is controlled by antibiotics, which influenced how the bacteria could move around in the medium that constituted the ‘memory cell,’ in the case of this project. The HB101 was able to approach the reader as it was surrounded by streptomycin-laced agar, and these cells (not the Nova Blue ones) were resistant to this antibiotic.
Potential Disadvantages of Living Data Storage
Antibiotics also motivated the HB101 to pick up the data-loaded plasmids as they also contained the data on tetracycline resistance. This antibiotic was present in the medium located between the data ‘storage’ and ‘reading’ areas. Therefore, the HB101 cells needed tetracycline resistance to reach the reading device.
This seemed like an elegant way to keep each cell type where they need to be in the bacterial network, which the researchers intended to set up to manage their data.
However, the risks implied by this reliance on antibiotics are clear. For example, someone with malicious intent could presumably wipe a hypothetical DNA-based archive with a completely different form of antibiotic.
In addition, and as with many forms of emerging technology, the desired process in the case of this project was reported to be relatively slow. HB101 cells took a reported approximation of 72 hours to reach the data-reader following plasmid pick-up. Therefore, DNA-based data transfer will need to get much faster for any future viability.
However, the project did achieve its main goal, which was to demonstrate that the reuse of a DNA data archive is possible. So, humanity may have an alternative to the resource-hungry necessity that modern inorganic data storage has become.
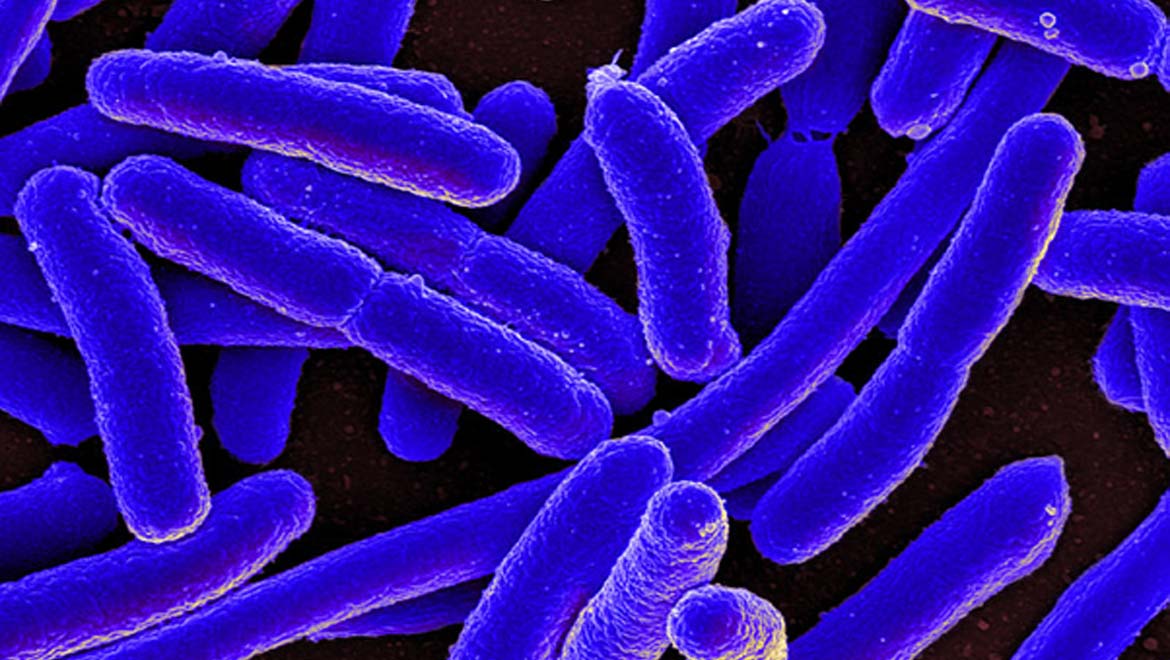
Top Image: The data in this study was stored in the DNA of E. coli bacterial colonies. (Source: NIAID - E. coli bacteria)
References
F. Tavella, et al. (2018) DNA Molecular Storage System: Transferring Digitally Encoded Information through Bacterial Nanonetworks. arXiv.org. arXiv:1801.04774v2.
Storing data in DNA is a lot easier than getting it back out, 2018, MIT Technology Review, https://www.technologyreview.com/s/610071/storing-data-in-dna-is-a-lot-easier-than-getting-it-back-out/ (accessed on 24 May 2018)
Waterford researchers develop new method to store data in DNA, 2018, rte.ie, https://www.rte.ie/news/ireland/2018/0219/941956-dna-data/ (accessed on 24 May 2018)






No comment