New approaches to studying biological problems have been sprouting up like mushrooms. Genomics is the study of all the genes of an organism. Proteomics is the study of all the proteins of an organism. Transcriptomics is the study of all the RNA molecules made by an organism. Collectively, these new approaches are known as “omics.” They are novel in that they use techniques and computational biology software that can track thousands of molecules at once. In the past, scientists could only track a maximum of a few genes at a time.
The research group led by Giulio Superti-Furga has been using all of these omics, and more, in their study of the immune response. The omics have identified molecules that might have critical roles in the body’s fight against viruses. Their importance has been confirmed using another hot new tool, CRISPR.
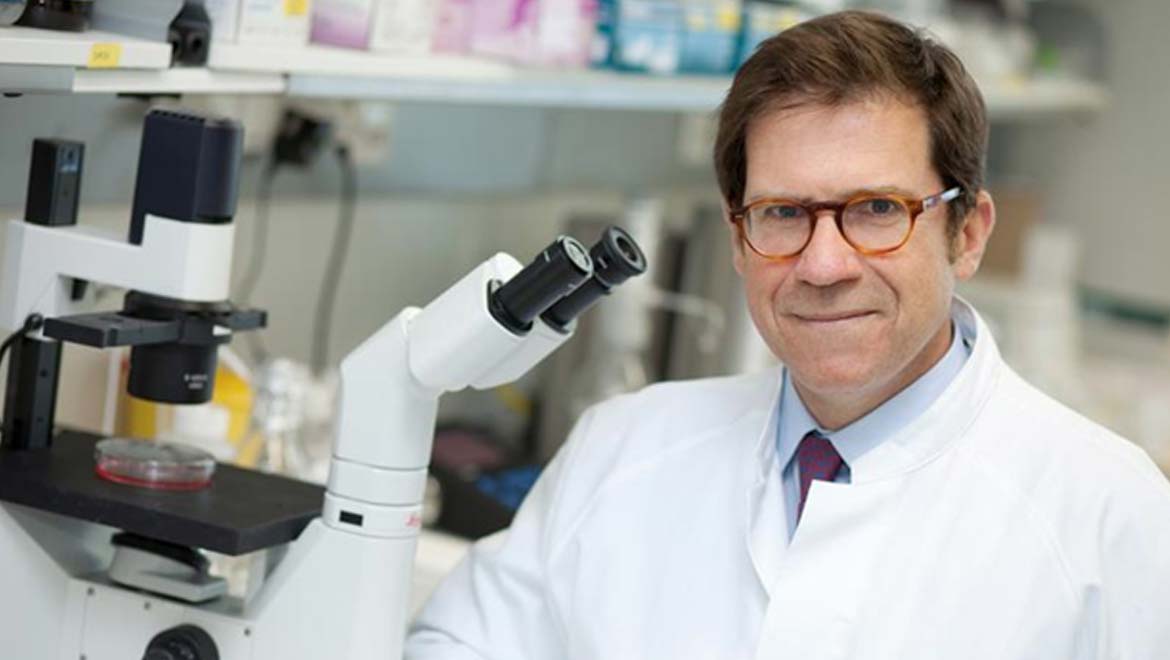
Professor Superti-Furga was born and educated in Milan and now is based in Vienna at the Research Center of Molecular Medicine. His group examined how cells in the immune system, the bodies defense against foreign invaders, gear-up to fight a viral infection. They did their work in mouse immune cells grown in culture.
Why omics?
The need for omics arises from two foundational concepts of modern biology. The first is the “Central Dogma of Molecular Biology” put forward by Nobel Laureate Francis Crick in 1958. This is the idea that, in cells, DNA genes provide the information to make slightly different versions of themselves, known as RNA transcripts. These RNA transcripts, in turn, are used to make proteins. Proteins do the vast majority of the work that defines the behavior of the cell. So, genes work through the RNA transcripts and functional proteins they produce.
The second foundational concept is “Genomic Equivalence.” This is the idea that in living things composed of many cells, each cell has all the genes used by every cell. Although there are a few exceptions to this rule, it is why the cloning of animals works. The first cloned sheep was made from the DNA of a breast cell in 1996. This proved that the breast cell had all the genes needed to make a complete sheep, not just the breast.
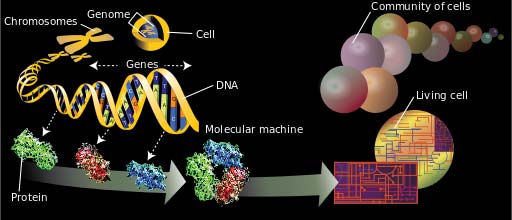
Diagram illustrating genomics. (Public Domain)
Well, you may ask, if every cell has the same DNA and genes, how can different kinds of cells in the body behave differently? And how can they change their behavior over time? Don’t genes control cells? The answer is: not every DNA gene is making its RNA transcript and its protein all the time in every cell. To really understand the differences between cells, it doesn’t really help to look at genes. It makes more sense to look at the collection of RNA transcripts they make, or the transcriptome. You can do this with transcriptomics.
Better yet, why not directly examine the collection of protein workhorses the cell is producing at any one time, the proteome? You do this with proteomics. The Superti-Furga group even went one step further, they examined the “phosphoproteome,” that is the proteins chemically modified with a phosphate group. You do with this with (wait for it) “phosphoproteomics.”
Remember, in the Central Dogma, not only must the protein be present, it must be functional. Some proteins only become functional when they receive phosphate groups at specific locations. Others only become functional when they lose phosphate groups at specific locations. Like a computer uses electrical circuits to process information, a cell can process information and make decisions using circuits composed of proteins gaining and losing phosphate groups.
OMG that’s a lot of omics!
Each omic has its own techniques, depending on the chemistry of the molecules they study. Of course, they complement each other, with the information gained using one set of omic techniques frequently suggesting experiments to perform using other omics. They all depend on the ability to keep track of hundreds or thousands of molecules at once with computational biology software. You must be able to compare the changing levels of specific molecules over time.
Suspects Identified
In the study by the Superti-Furga group, they infected immune cells grown in culture with viruses. Their results indicated that the addition of a phosphate to one particular protein, “MAVS,” was a prominent feature of the immune response. MAVS stands for “Mitochondrial Anti-Viral Signaling.” The mitochondrion is a part of the cell where it lives. They were also able to identify genes and proteins that seemed to be regulated in coordination with MAVS.
CRISPR Proof
When a DNA gene and its RNA transcript and protein are suspected of playing a critical role in a process, one way of proving they are truly necessary is to eliminate them. As a result of the logic of the Central Dogma, when you eliminate the gene in the DNA you also eliminate the corresponding RNA transcript and protein. When they are all gone, you then examine what function is missing. What is it the cell or organism is no longer able to do? Whatever is missing, is what that gene/transcript/protein does.
The Superti-Furga group used a technique to eliminate suspect genes that is going to dramatically affect all our lives, CRISPR. CRISPR will become powerful because it is so easy to use and so precise in its ability to eliminate genes. It can also edit genes. Its potential is not limited to the laboratory. It might be used in medical techniques or to eliminate genes and even organisms in the wild.
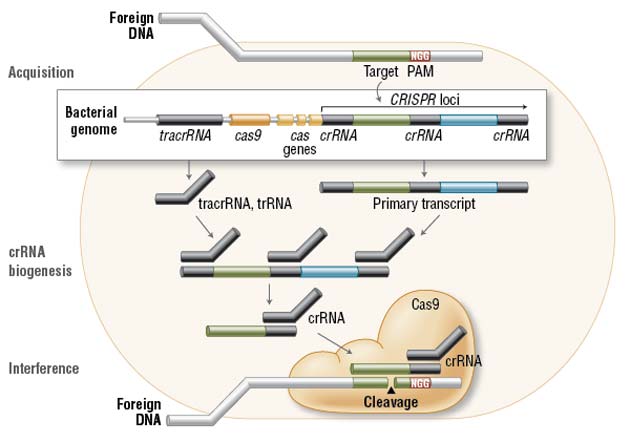
In the acquisition phase, foreign DNA is incorporated into the bacterial genome at the CRISPR loci. CRISPR loci is then transcribed and processed into crRNA during crRNA biogenesis. During interference, Cas9 endonuclease complexed with a crRNA and separate tracrRNA cleaves foreign DNA containing a 20-nucleotide crRNA complementary sequence adjacent to the PAM sequence. (Figure not drawn to scale.) (NEB)
CRISPR stands for “Clustered Regularly Interspersed Palindromic Repeats.” It is an immune system from bacteria that helps them fight off viral infections. When some species of bacteria survive a viral attack, they keep a copy of a fragment that virus’ genes as a short palindromic (back to front) repeat in their own DNA. When they collect a bunch of fragments it becomes a cluster that they pass on to their offspring. Bacteria refer to this cluster of sequences, like a contacts list on your phone, and when they encounter the sequences again they use enzymes to destroy them. Scientists can incorporate the CRISPR system in a wide variety of cells and manipulate it to destroy the sequence of the scientists’ choice.
When the Superti-Furga group turned this weapon on genes responding to the viral infection in the mouse cells, they were able to show that part of the response was eliminated as the gene was eliminated.
Bringing it all together
Giulio Superti-Furga and co-workers are trying to tease apart the changes in cell chemistry that occur after viral infection. After exhaustively cataloging changes in the transcriptome, proteome, and phosphoproteome they were able to identify key players in the process. In addition, they were able to prove the key role some genes play by eliminating them using CRISPR.
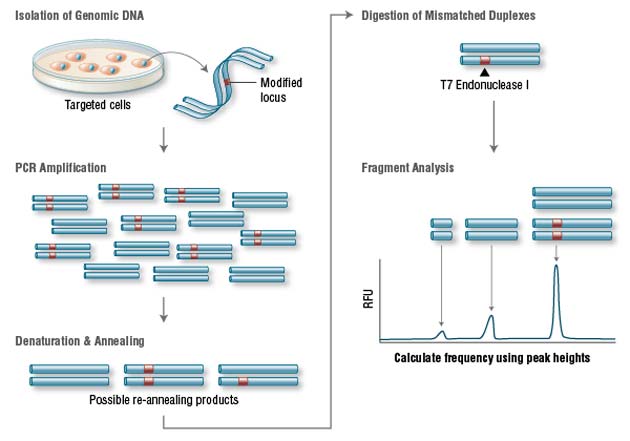
Genomic DNA is amplified with primers bracketing the modified locus. PCR products are then denatured and re-annealed yielding 3 possible structures. Duplexes containing a mismatch are digested by T7 Endonuclease I. The DNA is then electrophoretically separated and fragment analysis is used to calculate targeting efficiency. (NEB)
Scientists are now applying these techniques to all sorts of problems. Rather than examining the function of a few molecules at a time, they are looking at all the molecules, all the time. This is expected to greatly accelerate the pace of discovery.
Top image: Giulio Superti-Furga in CeMM laboratory (Meduniwien)
References
Richard K Kandasamy, Gregory I Vladimer, Berend Snijder, André C Müller, Manuele Rebsamen, Johannes W Bigenzahn, Anna Moskovskich, Monika Sabler, Adrijana Stefanovic, Stefania Scorzoni, Manuela Brückner, Thomas Penz, Ciara Cleary, Robert Kralovics, Jacques Colinge, Keiryn L Bennett & Giulio Superti-Furga, 2016, A time-resolved molecular map of the macrophage response to VSV infection
Available at: https://www.nature.com/articles/npjsba201627
Giulio Superti-Furga Bio, 2017,
Available at: http://superti-furga-lab.at/giulio-superti-furga-28.html
Shelly L. Ward, 2014, OMICS, BIOINFORMATICS, COMPUTATIONAL BIOLOGY
Available at: http://alttox.org/mapp/emerging-technologies/omics-bioinformatics-computational-biology/
Alex Reis, Ph.D., Breton Hornblower, Ph.D., Brett Robb, Ph.D. and George Tzertzinis, Ph.D., 2014, CRISPR/Cas9 and Targeted Genome Editing: A New Era in Molecular Biology






No comment